|
Generative Enriched Sequential Learning (ESL) Approach for Molecular Design via Augmented Domain Knowledge
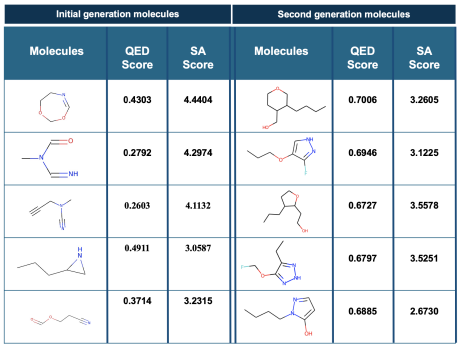
M. S. Ghaemi, K. Grantham, I. Tamblyn, Y. Li, H.K. Ooi Canadian AI (2022) Generative machine learning models for molecular design often struggle to produce molecules with desirable chemical properties because they focus primarily on learning the statistical distribution of molecular structures without incorporating domain-specific knowledge. In this work, we introduce a Generative Enriched Sequential Learning (ESL) approach that augments traditional sequential learning models -- including Hidden Markov Models, Recurrent Neural Networks, and Long Short-Term Memory networks -- with domain knowledge such as quantitative estimates of drug-likeness scores (QEDs). By incorporating these supervised chemical property metrics directly into the training process, the ESL method enables the model to learn specific patterns of particular interest more effectively, leading to the generation of de novo molecules with improved drug-likeness properties. This approach addresses the bias that standard generative models have toward prevalent molecules in training data, instead steering the generation process toward molecules with greater practical relevance for drug discovery. |